Lesson 3. Data Workflow Best Practices - Things to Consider When Processing Data
Learning Objectives
- List best practices for implementing data workflows.
- Be able to identify aspects of a workflow that can be modularized (i.e. ideal for functions) and be tested.
Data Workflow Best Practices
In the previous lessons, you designed a workflow to process several sites worth of Landsat data, but of course, identifying the workflow is just beginning.
For each of the steps you identified in your pseudocode, you need to flesh out how you can accomplish each task.
Before you design your workflow in more detail, there are many things to keep in mind regarding how you will use and name files, how you will modularize code to reduce repetitive code, etc.
Consider the data workflow best practices discussed below.
Do Not Modify the Raw Data
Typically you do not want to modify the raw data, so that you can always reproduce your output from the same inputs. In some cases, this is less important, e.g., when you are not tasked with curating the raw data. For example you could redownload the landsat data provided in this class.
In other cases, this is extremely important - you wouldn’t want to recollect a summers worth of field collected data because you accidentally modified the original spreadsheet.
A good rule of thumb is to create an “outputs” directory where you store outputs. An outputs directory is helpful because:
- You can always delete and recreate the outputs without worrying about deleting your original data.
- An outputs directory is expressive and reminds anyone looking at your project (including you in the future - i.e. your future self!) where the output products are - they don’t have to dig for the data.
Create Expressive Intermediate File Names and Outputs
You want to carefully consider how you name your intermediate and end product outputs so that the names reflect your intention and the content. For instance - in this case you are creating an NDVI dataframe which would be exported to a csv.
Here are some naming options:
# Less than ideal name
data.csv
# OK Name
ndvi.csv
# More expressive name
all-sites-ndvi.csv
Always use clear, expressive names for objects, files, etc. Ask yourself if someone reading your code could guess what is contained in that object or file based on the name.
Write Modular Code
As you are working on your pseudo code, consider the parts of your analysis that are well suited for functions.
Writing custom functions for repetitive tasks in your workflows can help your workflow to be more efficient and easier to follow and implement.
Ideally, a Python function will have 1-3 inputs and will produce 1-2 outputs.
Keep functions small and focused. A function should do one thing well.
You can then combine many functions into a wrapper function to automate and make for a nicely crafted program.
Functions - What Are They Again?
A function is a unit of code that executes a specific, outlined task on command, using sets of input parameters to specify how the task is performed.
input parameter(s) –> function does something –> output result(s)
Useful for organizing and executing generalizable code that you need to run frequently:
- importing data
- aggregating or summarizing data
- calculating indices such as normalized difference vegetation index, NDVI
- plotting data
You can use existing functions from commonly used packages (i.e. geopandas.read_file() or rasterio.open()) or write your own functions when one does not already exist.
Review of Key Components of Python Functions
defkeyword- Function name
- Input parameters/arguments
- Docstring (that identifies parameters and returns)
returnstatement
# Example of a simple function
def add_five(x):
"""Add the numeric value 5 to an input value.
Parameters
----------
x : int or float
Returns
----------
int or float
Input data with value increased by 5.
"""
return (x + 5)
# How do we call a Python function again?
add_five(3)
8
Now, check out the more complex geopandas.read_file() function on GitHub and identify each of the key components.
Do we have to fully understand all of the detailed code in order to use the function?
Simply put - no. We only need to know the appropriate inputs and outputs, so we can provide the correct file to get back our GeoDataframe.
This one of the primary benefits of writing and using functions!
Review of Benefits of Using and Writing Functions
- Reusability (e.g. by yourself or others)
- Fewer variables (e.g. temporary variables not needed outside of the function are not stored)
- Documentation/Reproducibility (e.g. for yourself or others)
- Easier updates to code (i.e. update only the function definition)
- Testing (e.g. include checks and tests on inputs and outputs directly within function)
- Modularity (i.e. stand-alone units of code that can executed independently and asynchronously)
Sometimes you may need to write custom functions to:
- complete tasks that do not already have published functions
- combine existing functions into one function for a specific task
Generalizing Custom Functions
The best functions complete one specific task but are generalizable for more than one application.
Example: Goal is a function that can be used to calculate Normalized Difference Vegetation Index (NDVI)
(Near_Infrared - Red) / (Near_Infrared + Red)
How Can We Generalize This For Broader Use?
There are other indices that can be calculated using the same formula:
# Normalized Burn Ratio (NBR)
(near_infrared - shortwave_infrared) / (near_infrared + shortwave_infrared)
# Normalized Difference Water Index (NDWI)
(green - near_infrared) / (Green + near_infrared)
For example:
(band1 - band2) / (band1 + band2)
Even better for conciseness and readibility:
(b1 - b2) / (b1 + b2)
def normalized_diff(b1, b2):
"""Calculate the normalized difference of two arrays of same shape.
Math will be calculated (b1-b2) / (b1+b2).
Parameters
----------
b1, b2 : numpy arrays
Two numpy arrays of same shape.
Returns
----------
n_diff : numpy array
The element-wise result of (b1-b2) / (b1+b2) calculation.
"""
n_diff = (b1 - b2) / (b1 + b2)
return n_diff
Expand Code to Add Checks and Tests
By including checks (i.e. tests) in your code, you can be more confident that your code is doing what you think is!
Specific Reasons to Check Your Code
- Check that inputs are of correct type/format (e.g. both arrays are two-dimensional)
- Check that necessary preqrequisites have been executed or exist (e.g. a directory named
output) - Test assumptions of code (e.g. is it actually doing what you think it is?)
- Identify points of failure (e.g. where is the code failing - input data, processing/analysis, writing out data?)
- Identify something about the function that you did not consider (e.g. function is applicable to additional data types, function needs additional code to handle special circumstances)
How Can You Check Your Code?
There are many different ways to start checking your code. You can begin with conditional statements and work toward more complex checks using try/except and exception handling (which are introduced below).
Conditional Statements
This is frequently referred to as “asking permission” before code can execute.
if condition_1:
action_1
elif condition_2:
action_2
else:
action_3
Thinking about the norm_diff function: what is a condition that we might want to check for the code to run successfully?
Take a look at the equation again, and think about what the intended inputs are for b1 and b2 (“arrays of same shape”).
(b1 - b2) / (b1 + b2)
import numpy as np
# Create numpy array inputs for function
nir_band = np.array([[6, 7, 8, 9, 10], [16, 17, 18, 19, 20]])
red_band = np.array([[1, 2, 3, 4, 5], [11, 12, 13, 14, 15]])
# Create three dimensional numpy array for testing
nir_band_3d = np.array([[[6, 7, 8, 9, 10], [16, 17, 18, 19, 20]]])
nir_band_3d.ndim
3
# Note that the if/else is used when calling the function
if nir_band_3d.shape == red_band.shape:
ndvi = normalized_diff(b1=nir_band_3d, b2=red_band)
print(ndvi)
else:
print("Input arrays are not of same shape")
Input arrays are not of same shape
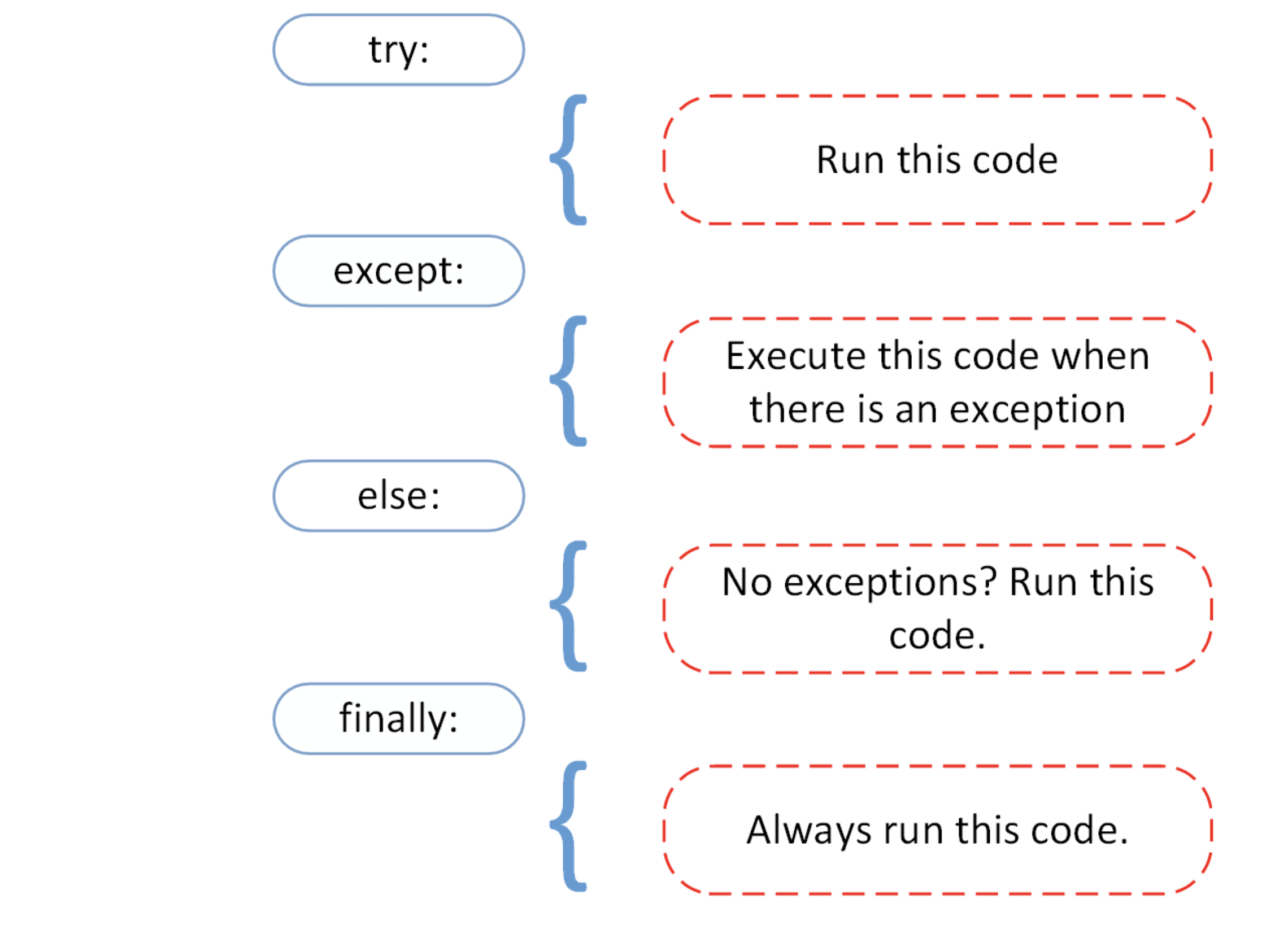
Try and except
These statements allow the a code block to try to execute first, and then do something else if the code is not executed successfully.
This is often referred to as “asking for forgiveness, rather than permission”.
# Example of asking for permission
x = 5
y = 0
if y != 0:
print(x/y)
else:
print("Division by zero not allowed!")
Division by zero not allowed!
# Instead, ask for forgiveness
try:
print(x/y)
except:
print("Division by zero not allowed!")
Division by zero not allowed!
# In this example, try/except is used when calling the function
try:
ndvi = calc_indices.norm_diff(b1=nir_band_3d, b2=red_band)
print(ndvi)
except:
print("Input arrays are not of same shape")
Input arrays are not of same shape
After this try/except, if you try to print ndvi, you will see the following error because it was not created by the code.
---------------------------------------------------------------------------
NameError Traceback (most recent call last)
<ipython-input-8-24938b8c9bd5> in <module>
6 print("Input arrays are not of same shape")
7
----> 8 print(ndvi.ndim)
NameError: name 'ndvi' is not defined
Caveat: Do you see anything potential issues with try and except statements?
import os
import earthpy as et
# Another example of try and except
directory = os.path.join(et.io.HOME, "earth-analytics", "test")
# Create folder if it does not exist
try:
os.makedirs(directory)
print("Directory created!")
except:
print("Directory already exists!")
Directory created!
Caveat: How would you know that a failure to make a new directory was due to it already existing?
Is it possible that something else could cause a new directory to not get created?
Exception Handling
Also commonly referred to as error handling, exceptions are used to check for specific types of errors that can occur when running code.
Though it is more common to use exception handling when writing code that you want to publish as packages or libraries or when you are writing applications for end users, it is useful to know about these, so that you are familiar with them when you receive exception messages by others’ code.
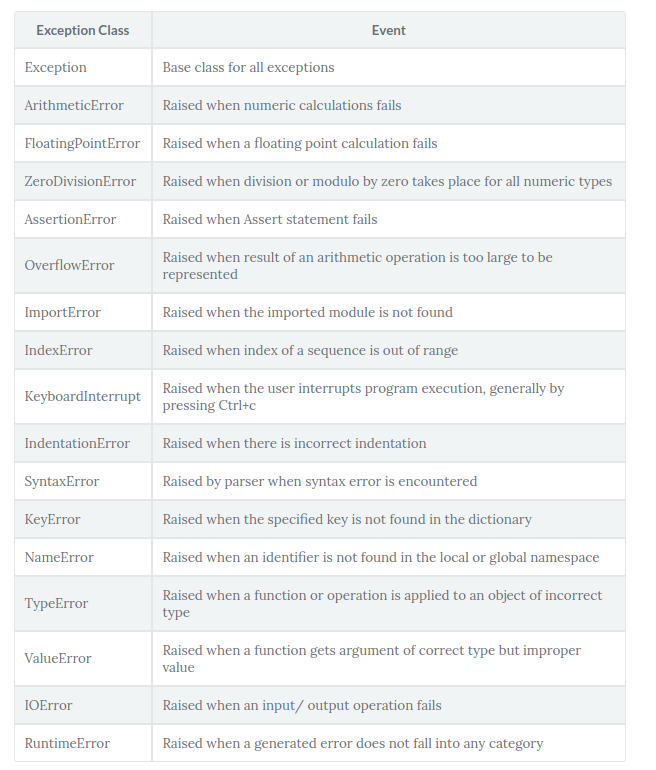
Commonly Used Exceptions
# Same try and except with exception FileExistsError
directory = os.path.join(et.io.HOME, "earth-analytics", "test")
try:
os.makedirs(directory)
print("Directory created!")
except FileExistsError:
print("Directory already exists!")
# Uncomment line below to have the except do nothing (e.g. no print statement)
#pass
Directory already exists!
# You could also explore error handling in your function definitions
# Example of ValueError
def normalized_diff(b1, b2):
"""Calculate the normalized difference of two arrays of same shape.
Math will be calculated (b1-b2) / (b1+b2).
Parameters
----------
b1, b2 : numpy arrays
Two numpy arrays of same shape.
Returns
----------
n_diff : numpy array
The element-wise result of (b1-b2) / (b1+b2) calculation.
"""
if not (b1.shape == b2.shape):
raise ValueError("Inputs arrays should have the same dimensions")
n_diff = (b1 - b2) / (b1 + b2)
return n_diff
Now if you try to run the following code:
ndvi = normalized_diff(b1=nir_band_3d, b2=red_band)
You will see the following ValueError that can help you (and others) understand why the function did not execute successfully.
---------------------------------------------------------------------------
ValueError Traceback (most recent call last)
<ipython-input-13-7a66955ecfea> in <module>
----> 1 ndvi = normalized_diff(b1=nir_band_3d, b2=red_band)
2
<ipython-input-11-018feb34e9f1> in norm_diff(b1, b2)
16 """
17 if not (b1.shape == b2.shape):
---> 18 raise ValueError("Inputs arrays should have the same dimensions")
19
20 n_diff = (b1 - b2) / (b1 + b2)
ValueError: Inputs arrays should have the same dimensions
Old Fashioned Trial and Error
Maybe the code is executing just fine, but maybe you are not getting the output in the most optimal format.
It may not be clear to you until you run the function many times with different inputs.
# Create numpy array inputs for function
# Note that we are creating a zero in the numerator with -15
nir_band = np.array([[6, 7, 8, 9, 10], [16, 17, 18, 19, -15]])
red_band = np.array([[1, 2, 3, 4, 5], [11, 12, 13, 14, 15]])
# Produces infinity values due to divide by zero
ndvi = normalized_diff(b1=nir_band, b2=red_band)
print(ndvi)
[[0.71428571 0.55555556 0.45454545 0.38461538 0.33333333]
[0.18518519 0.17241379 0.16129032 0.15151515 -inf]]
<ipython-input-12-f04f35d1b23a>:21: RuntimeWarning: divide by zero encountered in true_divide
n_diff = (b1 - b2) / (b1 + b2)
What if we would rather have the output be a masked numpy array if there are any infinite values or nan values?
# Function definition in the earthpy package:
# https://github.com/earthlab/earthpy/blob/master/earthpy/spatial.py#L62
import warnings
def normalized_diff(b1, b2):
"""Take two numpy arrays and calculate the normalized difference.
Math will be calculated (b1-b2) / (b1+b2). The arrays must be of the
same shape.
Parameters
----------
b1, b2 : numpy arrays
Two numpy arrays that will be used to calculate the normalized difference.
Math will be calculated (b1-b2) / (b1+b2).
Returns
----------
n_diff : numpy array
The element-wise result of (b1-b2) / (b1+b2) calculation. Inf values are set
to nan. Array returned as masked if result includes nan values.
"""
if not (b1.shape == b2.shape):
raise ValueError("Both arrays should have the same dimensions")
n_diff = (b1 - b2) / (b1 + b2)
# Set inf values to nan and provide custom warning
if np.isinf(n_diff).any():
warnings.warn(
"Divide by zero produced infinity values that will be replaced with nan values",
Warning)
n_diff[np.isinf(n_diff)] = np.nan
# Mask invalid values
if np.isnan(n_diff).any():
n_diff = np.ma.masked_invalid(n_diff)
return n_diff
ndvi = normalized_diff(b1=nir_band, b2=red_band)
print(ndvi)
[[0.7142857142857143 0.5555555555555556 0.45454545454545453
0.38461538461538464 0.3333333333333333]
[0.18518518518518517 0.1724137931034483 0.16129032258064516
0.15151515151515152 --]]
<ipython-input-15-6acb522572a0>:25: RuntimeWarning: divide by zero encountered in true_divide
n_diff = (b1 - b2) / (b1 + b2)
<ipython-input-15-6acb522572a0>:29: Warning: Divide by zero produced infinity values that will be replaced with nan values
warnings.warn(
There is now a warning provided by the function to explain that the division by zero resulted in values that are replaced with nan values.
There are many ways to include tests and checks in your code, so explore the resources below to learn different ways to incorporate them into your workflows.
Additional Resources
Share on
Twitter Facebook Google+ LinkedIn
Leave a Comment